These Matlab programs have recently been uploaded to my Inria gitlab repository. The first one has been circulated “informally”; the next ones have been used to illustrate articles.
Sequential Monte Carlo (SMC)
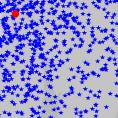
I am quite happy to propose a rather nice series of demonstration software for particle filtering, also called Sequential Monte Carlo (SMC), for applications in tracking. SMC demos is written in matlab.
View page or gitlab repository.
Individual-based models (IBM)
I have also developed several Individual-based models (IBM): they are mathematical models that is used to study the behavior of systems made up of many individual components, such as a population of animals or a network of interconnected nodes. In an IBM, each individual is represented as a separate entity, with its own set of characteristics and behaviors. The behavior of the system as a whole is determined by the interactions among these individuals, as well as by external factors such as the environment. IBMs are useful because they allow us to study the behavior of complex systems in a more detailed and realistic way than is possible with traditional aggregate models. They are often used in ecology, epidemiology, and other fields where the behavior of individual entities is important.
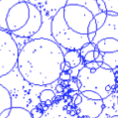 Forest IBM is an individual-based model of forest dynamics. It is a simple extension of the model proposed by Bolker and Pacala that also includes an extra component representing the radius of the zone of influence.
Forest IBM is an individual-based model of forest dynamics. It is a simple extension of the model proposed by Bolker and Pacala that also includes an extra component representing the radius of the zone of influence.
View page and gitlab repository.
 IBM cellulose is an individual-based model for the degradation of one cellulose bead (dozens of micrometers in diameter) by cellulolytic bacteria .
IBM cellulose is an individual-based model for the degradation of one cellulose bead (dozens of micrometers in diameter) by cellulolytic bacteria .
View page and gitlab repository.
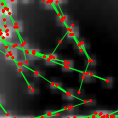 IBM clonal: An Individual-based model simulator for clonal plant growth developed in 2011 and 2012 in the context of the ANR Syscom project Modecol, in Matlab.
IBM clonal: An Individual-based model simulator for clonal plant growth developed in 2011 and 2012 in the context of the ANR Syscom project Modecol, in Matlab.
View page and gitlab repository.
