
Algorithms in Structural Bio-informatics :
Winter School
2-7 december 2012, Inria Sophia Antipolis, France

Conference guide
Agenda
The first afternoon (Dec. the 2nd) will be devoted to short presentations. Each of the five remaining days will be organized as follows:
- Lecture from 9am to 10:30am, and from 10:45 am to noon
- Lunch at 12:45pm (sharp!)
- Practical from 2pm to 5:30pm
[Sunday the 2nd, 2pm - 6:30pm] Short presentations by the participants (7'+3')
[Monday the 3rd] I. Obtaining and organizing structural information for modeling studies
Instructors:
- Joël Janin, IBBMC, Orsay
- Charles H. Robert, Laboratory of Theoretical Chemistry, CNRS

Topics: Projects proposed:
- The Protein Data Bank and related resources
- Organizing and manipulating protein structural data and derived quantities
- Data structures for higher-level problems
- Analyzing conformational changes in protein-protein association
- Correlating observed structural changes and collective movements of proteins
[Tuesday the 4th] II. Modeling protein complexes and assembies with Voronoï diagrams
Instructor:
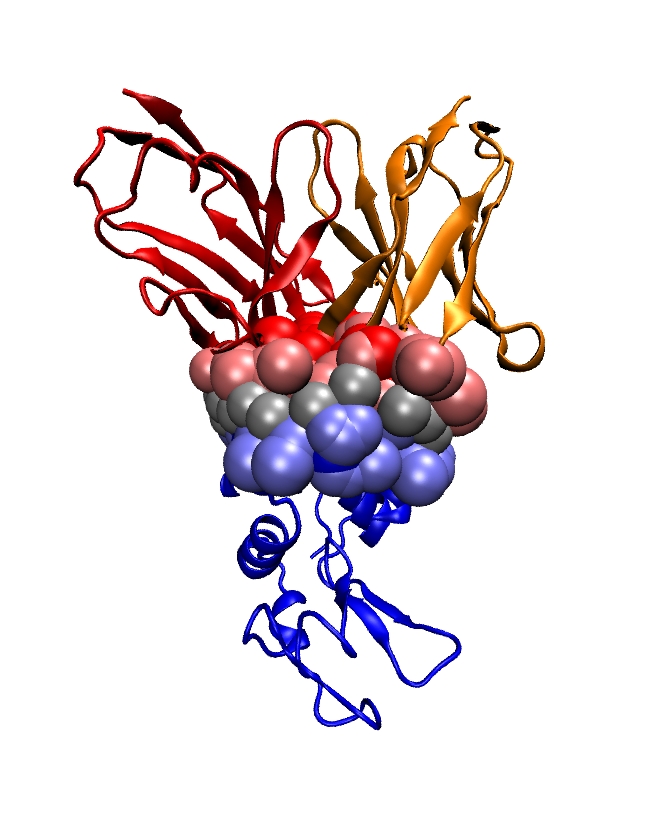
Topics: Projects proposed:
- Affine and curved Voronoi diagrams: a primer.
- Modeling atomic resolution protein interfaces and binding patches.
- Modeling fuzzy protein assemblies in the context of reconstruction by data integration.
- Investigating complexes from the immune system, annotated in the IMGT/3Dstructure-DataBase
- Spotting putative binding patches on orphan proteins.
[Wednesday the 5th] III. Molecules as robots: mining the flexibility of proteins
Instructor:
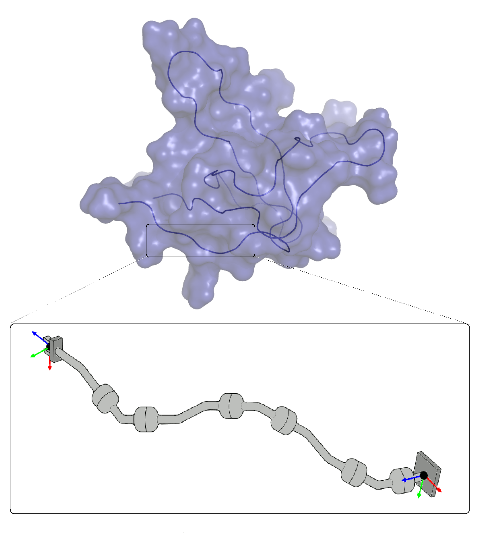
Topics: Projects proposed:
- Some basic notions on robot modeling and motion planning algorithms.
- Robotics-inspired methods to explore the conformational space of proteins.
- Investigating the accessibility to active sites considering flexible molecule models.
- Computing conformational transitions of proteins.
- Simulating ligand (un)binding with a single processor.
[Thursday the 6th] IV. Structural comparisons
Instructors:
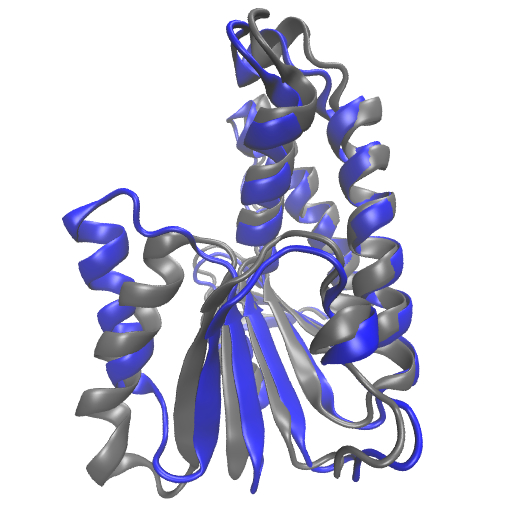
Topics: Projects proposed:
- Integer programming modeling and relaxations
- Application to the study of protein structures
- Application to the study of protein binding patches
- Studying protein flexibility using the Comparative Structural Alignment web-server
- Identifying orphan protein structures
[Friday the 7th] V. Docking algorithms
Instructors:
- Chantal Prévost, Laboratory of Theoretical Chemistry, CNRS
- Martin Zacharias, Physik-Department, Technische Universität München
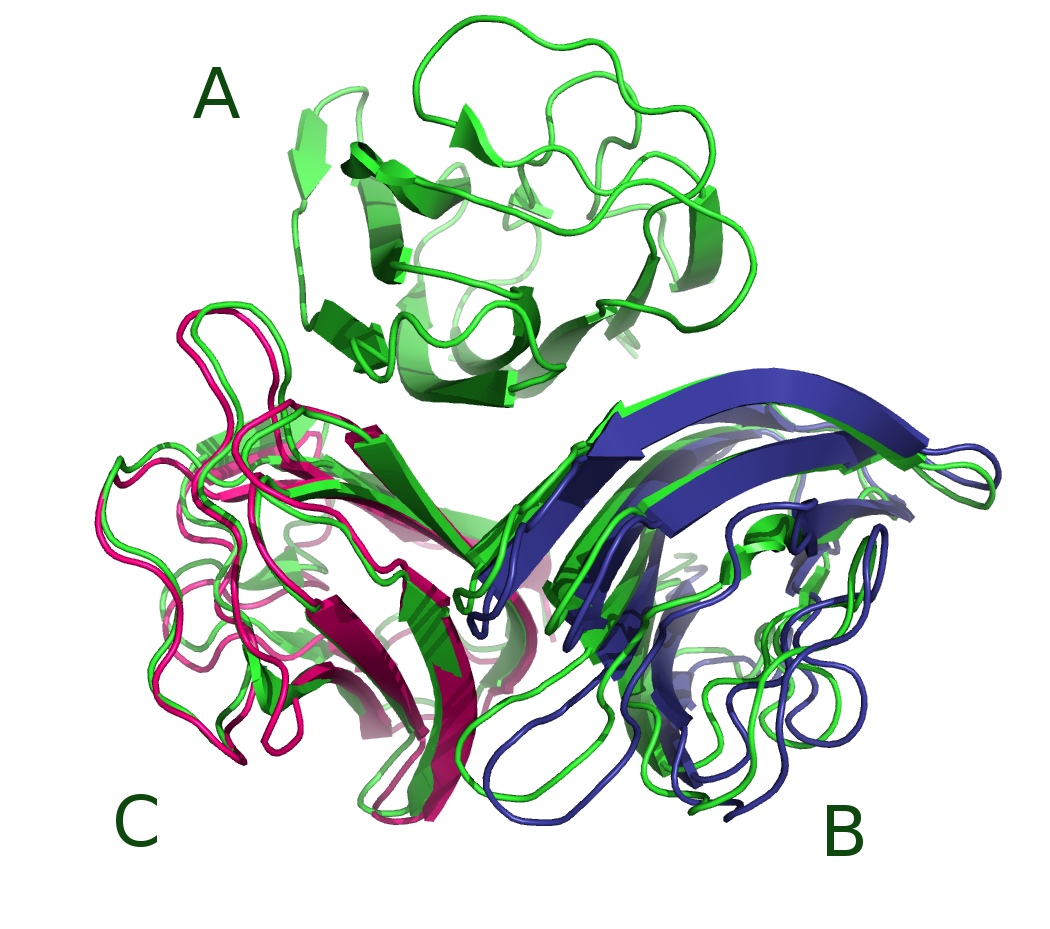
Topics: Project proposed:
- Macromolecular docking methods: principles and state of the art
- Coarse-grained representations of proteins and nucleic acids
- The ATTRACT/PTools docking suite
- Systematic docking including side chain and global flexibility
- Docking multiple partners
- Construction of protein filaments: from two-component docking to the generation of full helices
| webmaster |