model invariant by rotations and translations
As for the first step, missing catorgaphic data are modelled
by a null label![]() .
.
Second order potentials
The potentials on the cliques of order 2 induce some constraints on the consistency of angles between neighboring chains and the corresponding map segments :
1) two chains matched to the same segments should make an angle close to PI.
2) two connected chains matched to two connected segments should have an angle close to the corresponding segments.
3) two connected chains matched to two non connected segments
are penalized.
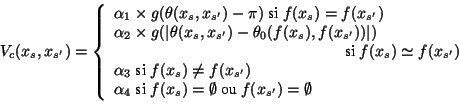
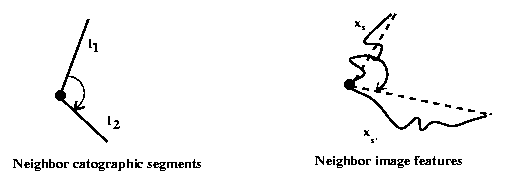
The function g is defined on ![]() ,
convex and minimum in 0 ; g drives the strength of the prior on angles.
We consider a parabolic function.
,
convex and minimum in 0 ; g drives the strength of the prior on angles.
We consider a parabolic function.
![]()
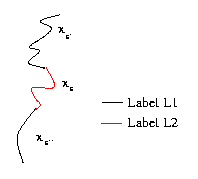
This last term adds some consistency constraints between the length of the chains and the corresponding segments.
It is defined using the features length:
![]()
This term induces non markovian properties as it
includes all sites of the models. However, we still have a Gibbs distribution.
The energy is minimized by a simulated annealing
using a Metropolis dynamics but we can also use an ICM because the first
step (pixel labelling) provides a good initialization. This last algorithm
is faster than the simulated annealing.